Traits Computation
The traits computation web service supports traits-based studies of phytoplankton communities. It computes demographic and morphological traits such as biovolume (µm³), surface area (µm²), surface/volume ratio, density (cells/L), cell carbon content (pg), total biovolume (µm³/L), total carbon content (pg/L).
Please, visit the Atlas of Shapes for association of each taxa of your dataset with the most similar geometric shape selected from the catalogue of proposed models.
Follow the operational steps to run the service. It will return an output file in .csv format, including all input data and the new calculated traits.
Input file must be in .csv format.
- dataset
- traits
- calculation type
Step 1 - upload your dataset or select a demo file (*)
Upload your dataset structured according to the Phytoplankton Data Template, or select a demo file from the list of datasets about phytoplankton harvested from the LifeWatch ERIC Metadata Catalogue (you can use the filter to find a specific dataset):
|
|
Step 2 - select the traits (*)
You can select one or more traits to be computed. Check the "Select all" option if you want to compute all available traits:
Step 3 - select the calculation type (*)
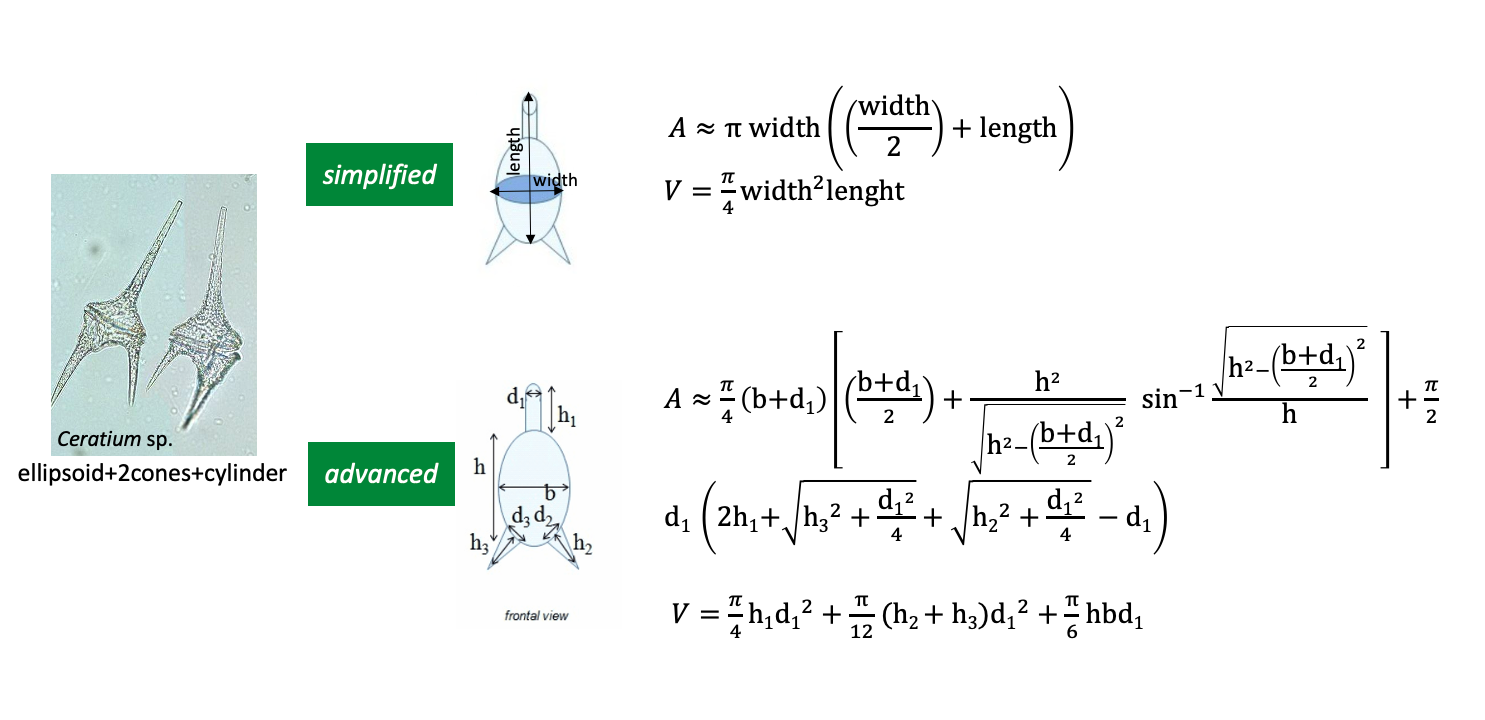
You can choose between two computation modalities for the biovolume and surface area traits:
- advanced: it provides a more accurate estimation of biovolume and surface area by using more detailed measurements for the linear dimensions in accordance to the shape and the view of the organism. The mandatory fields for this calculation type are: scientificname, measurementremarks and the measured linear dimensions.
- simplified: it provides an approximate estimation of biovolume and surface area based only on maximum and minimum linear dimensions (length and width). The mandatory fields for this calculation are: scientificname, length, width and measurementremarks (the sedimentation view of the organism e.g., frontal, lateral, unique, valvar and girdle);